Chemicals and instruments
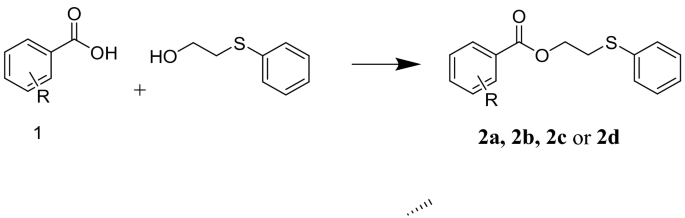
The following chemicals and biological materials were purchased from Frutarom (UK): β-carotene, DPPH, Na2HPO4/NaH2PO4, NaCl, gallic acid, trolox, methanol, starch, porcine pancreatic amylase, lipase, glucosidase, dinitrosalicylic acid (DNSA), Na2CO3, dimethyl sulfoxide (DMSO), and acarbose. The evaluated compounds 2a–2d were synthesized previously by our team, as outlined in Fig. 212.
The reaction of benzoic acid derivatives with thiophenyl ethanol. (When R = H presents compound 2a, R = 2-OH presents 2b, R = 3-OH presents 2c, and R = 4-OH presents 2d).
α-Amylase inhibitory activity
The α-amylase inhibitory evaluation was established following the Wickramaratne et al. procedure13 with minor changes. Working solutions with concentrations of 10, 50, 100, 500, and 1000 µg/ml were produced from our synthesized compounds, diluting them with a buffer of Na2HPO4/NaH2PO4 (0.02 M), NaCl (0.006 M) at pH 6.9 and then brined up to 10 ml using a 10 ml volumetric flask. The acarbose was considered a positive control and was established following the same previous steps utilized for the synthesized compounds. As percent inhibition, the enzyme inhibitory potential was expressed, and the following equation was utilized to estimate the IC50 dose for the samples14.
$$\%a-Amylase\ Inhibiton=\frac{Abs100\%\ control-Abs\ Sample}{Abs\ 100\%\ Control}\times 100$$
α-Glucosidase activity
The inhibition of α-glucosidase activity by the synthesized molecules was measured. The α-glucosidase inhibitory activity is expressed as percentages of inhibition, which was calculated using the following formula:
$$ {\text{Inhibitory}}\,{\text{effect }}\% \, = \, \left( {A_{b} – \, A_{s} } \right)/ \, A_{b} *100 \, \% $$
where Ab and As are the absorbance values of the blank (containing PBS, α-glucosidase, and PNPG, the colored substrate of glucosidase) and the tested samples containing PBS, samples, α-glycosidase, and PNPG, respectively.
The α-glucosidase activity was measured using 10 and 20 mg/ml of the synthesized molecules. Each concentration was recorded by using 1, 3, 6, 9, and 12 mm PNPG. The inhibitory pattern was assessed using a Lineweaver–Burk plot. The constant Ki of enzyme inhibitory effect was determined15.
Antioxidant activity
DPPH method
Four synthetic compounds were tested for the efficiency of scavenging free radicals matched with trolox and gallic acid as a basic. From the compounds, 1 mg/ml concentration solutions in methanol were prepared, and the solutions were used to prepare concentrations of 5, 10, 20, 30, 40, and 50 μg/ml. The DPPH reagent (0.002% w/v) was dissolved in methanol before mixed with working concentrations in ratios of 1:1:1 (compound:DPPH:methanol). The methanol solution was adopted as a blank. All the solutions were incubated for 30 min at room temperature in the dark. When the antioxidant compound reacts with DPPH, which can donate hydrogen, it is reduced the color changes from deep violet to light. Absorbance values were estimated by using a UV–Vis spectrophotometer at a wavelength of 517 nm. The DPPH inhibition percentage of all tested compound, trolox and gallic acid, was estimated according to the formula:
$$ {\text{Inhibition}}\,{\text{ percentage }} = \, \left( {A_{b} – A_{s} } \right)/A_{b} \times 100\% , $$
where Ab is blank absorbance, and AS is sample absorbance. The IC50 values for tested synthetic compounds, trolox and gallic acid were determined from the curves16.
β-Carotene method
The activity of the synthesized compounds was assessed via performing a modified assay of Gazzani and Miller17. The method was based on the coupled oxidation of β-carotene and linoleic acid emulsion. The bleaching mechanism of β-carotene is produced from the hydro-peroxides created from linoleic acid18. During the oxidation process, the characteristic orange color of β-carotene and the chromophore will be lost. The presence of antioxidants can hinder the extent of the β-carotene bleaching. Briefly, 1 mg of β-carotene was dissolved in 2 ml of chloroform, 20 mg of linoleic acid, and 200 mg of Tween 20. A rotary evaporator entirely vaporized chloroform at a temperature of less than 30 °C under reduced pressure. Then 200 ml of distilled water saturated with oxygen was added to the flask, which was shaken vigorously for 30 min. A sample (5 ml) of the prepared emulsion was transferred to a series of tubes containing 0.1 ml of the synthesized compounds or tocopherol (2 mg/ml).
The control sample was prepared in the same way but without added antioxidants. Each sample type was prepared in triplicate. The samples were inserted in a water bath at 50 °C for 2 h. The absorbance of the samples was measured using a spectrophotometer at a wavelength of 470 nm, immediately after the preparation of the samples, and at 15-min intervals until the end (t = 120 min) of the experiment.
Anti-lipase activity
The lipase inhibitory activity was conducted by following the method in the references with minor modifications19,20,21. The prepared synthesized compounds solutions (1 mg/ml) were diluted with 10% DMSO to produce five different dilutions (0.2, 0.4, 0.6, 0.8, and 1 mg/ml). Orlistat anti-obesity drug was considered a positive control in this protocol and was then tested following the same steps used previously. A Tris–HCl buffer was used to zero UV–Vis spectrophotometer at 405 nm. The % inhibition of the synthesized compounds and orlistat was calculated by using the following equation:
$$ {\text{Inhibitory}}\,{\text{lipase}}\,{\text{percentage}}\,\left( \% \right) = \left[ {\left( {A_{b} – \, A_{s} } \right)/A_{b} } \right]*100 $$
where Ab is blank absorbance, and AS is sample absorbance. The IC50 values for tested synthetic compounds and orlistat were determined from the constructed curves.
Molecular docking
The docking studies on the α-amylase enzyme were based on the human pancreatic amylase (the X-ray crystallographic structure (PDB code 4W93) that was obtained from the RCSB repository. This 3D protein structure was chosen based on its good resolution of crystallography (X-ray diffraction 1.352 Ao). Additionally, this work is considered a continuation of our previous work on the α-amylase enzyme that utilized the ID: 4W93 α-amylase crystal structure22.
Preparing this crude obtained crystal structure for docking, using the PyMOL program (v1.8), started with removing the native ligand, non-protein atoms, and the present crystallographic waters. This step was followed by protonating imidazole and amide side chains via an H++ server. Then the polar hydrogen atoms were added and the structure was brought up to a physiological pH. The open-source program rDOCK (rdock.sourceforge.net), a development of RiboDock, was applied to perform the docking studies23. Considering the cavity occupied by the native ligand (a flavonol glycoside called montbretin A) of the utilized protein crystal structure (code 4W93), the binding site for docking studies was defined within 6Ao around it. The default docking procedure was followed that includes 3 stages of Genetic Algorithm search (GA-3, GA-2, and GA-1). This was followed by low-temperature Monte Carlo (MC) and Simplex minimization (MIN) stages22.
In the current study, the most potent compound against α-amylase was the 2a compound. Applying the empirical score function of Rdock, the 2a molecule was chosen to be docked in the binding pocket of the prepared crystal structure, keeping twenty docking solutions for each inhibitor to be sorted by their binding scores and later visually investigated for the connections between the inhibitors and the pocket’s residues.
ADME-T calculations
A set of ADME-T related properties were calculated by using the QiKProp module (schrödinger 10.9, LLC, NY) running in normal mode. QikProp generates relevant descriptors and uses them to perform ADMET predictions24,25.
Statistical analysis
The antidiabetic, antioxidant, and antilipase experimental works were conducted in triplicates and the results were expressed as means ± SD standard deviation while the result was considered significant when the p value was < 0.05.